Protein-Protein Interactions
Despite many advances in the development of modeling, docking, and scoring methodology, correctly predicting protein-protein interactions remains challenging. In particular, selecting the final structure model(s) from all predictions is typically considered one of the most difficult steps and is often the most critical. We have developed a novel, physics-based, multi-step approach to identify near-native protein-protein complex structures from a set of top-ranked poses. In this method, steered molecular dynamics (MD) simulations are used to estimate the force required to separate the partners of docked protein-protein complexes by pulling one partner away from the other. For the top complexes (those with the highest force required for separation) more accurate but time-consuming free energy calculations (umbrella sampling combined with the weighted histogram analysis method (WHAM)) provide an estimate of the potential of mean force (PMF), and thus the free energy of complex dissociation. The method showed superior performance in identifying native protein-protein structures compared to any other tested method.
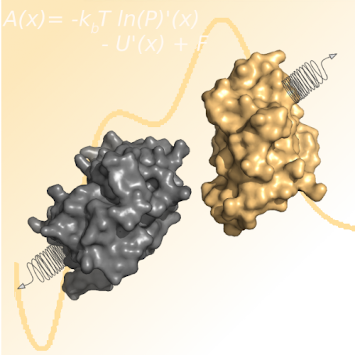
Kingsley, L.J.; Esquivel-Rodríguez, J.; Yang, Y.; Kihara, D.; Lill M.A. Ranking protein-protein docking results using steered molecular dynamics and potential of mean force calculations. J. Comput. Chem. 37, 2016, 1861-5
